Merry Christmas to everyone. As usual for this blog at this time of year, for your Christmas reading we will take a look at a particular aspect of human consumption, in this case alcohol.
James Bond was created in 1953 by Ian Fleming (who also created Chitty-Chitty-Bang-Bang, The Magical Car), and over a 14-year period there was a series of 12 novels and two short-story collections. The rights to the character were purchased for the film world in the 1960s, so that over the past 50 years we have had a franchise of 24 official films, plus two other licensed ones (Casino Royale in 1967, and Never Say Never Again in 1983).
Actually, the first licensed Bond film was a long-forgotten one made for CBS TV in 1954. This was a 1-hour version of Casino Royale, starring Barry Nelson as Bond, Peter Lorre as Le Chiffre, and Linda Christian as a renamed Vesper Lynd (see Barry Nelson - den bortglömde Bond).
This movie infographic (excluding the 2015 film, and the unofficial films) is from The Economist.
The Bond character
James Bond has been portrayed in films officially by six different actors, but the character remains essentially the same, although somewhat different from the one depicted in the books.
In early 1997, the monthly magazine Men's Health published an article in which doctors and psychologists commented on the life and lifestyle of the Bond character, the world's most un-secret secret agent (see Sprit, kvinnor och cigarretter tog livet av James Bond). The results were not good — Bond was either dead or close to it, as he was a paranoid, impotent alcoholic.
Bond's psychological profile was that of an emotionally stunted psychopath of type A who suffers from post-traumatic stress. According to Fleming's books, Bond was orphaned at age 11 (his parents died in a mountaineering accident), he lost his virginity in a brothel in Paris at 16, and killed his first mistress the following year. An ideal man to be a licensed assassin.
His massive daily alcohol consumption (all carefully documented in both the books and films) makes him a category 3 alcoholic. This means that he couldn't possibly have done his actual job competently; and it should also have led to violent temper outbursts (which may explain the government-sanctioned killing sprees). The liquor should also have led to a shrinking of his genitals, and have damaged his liver to the extent that it could no longer break down estrogen, so that he started to develop breasts and become impotent. His well-documented sexual excesses would also make him a prime candidate for sexually transmitted diseases. On top of this, the books (but not the films) also document a comprehensive smoking habit.
Bond was, of course, a form of wish-fulfillment for his creator, Ian Fleming, who was also a heavy drinker and smoker. He died of a heart attack at age 56, an age that Bond himself could not possibly have out-lived. Bond was more in danger from his own lifestyle than from SMERSH, or anyone else bent on world domination.
Bond is thus more a collection of memes than an actual character. This infographic is from the GBShowPlates website, and summarizes Bond's lifestyle.
The Bond drinks
Just about every aspect of Bond's career has been analyzed, and ranked, from the music to the cars to the watches, and most especially the women (the so-called "Bond girls"). However, much of the interest seems to lie in the booze, which is what we will look at here.
Along with coffee (and, once, tea), Bond has consumed copious amounts of alcohol, which he tends to drink alone, or in private settings. He is also what is known as a "label drinker", in that the brand is at least as important as the bottle's contents. This is a gift for the liquor industry, who, along with the car industry, are perpetually looking for opportunities for "brand placement" in films and sporting events. Fleming was chastised for introducing this into his books, but he simply replied that it was an attempt to round-out the character.
As far as the novels are concerned, they have received special medical attention by Graham Johnson, Indra Neil Guha, Patrick Davies (2013. Were James Bond’s drinks shaken because of alcohol induced tremor? British Medical Journal 347: f7255). They recorded every drink consumed in every book, calculated the number of alcohol units involved, and then converted that to daily intake (since the books are quite clear about their time span).
Their results are summarized in this infographic, from their article.
Basically, the medical results were as before:
Across 12 of the 14 books, 123.5 days were described, though Bond was unable to consume alcohol for 36 days because of external pressures (admission to hospital, incarceration, rehabilitation). During this time he was documented as consuming 1150.15 units of alcohol. Taking into account days when he was unable to drink, his average alcohol consumption was 92 units a week (1150 units over 87.5 days). Inclusion of the days incarcerated brings his consumption down to 65.2 units a week. His maximum daily consumption was 49.8 units (From Russia with Love day 3). He had 12.5 alcohol free days out of the 87.5 days on which he was able to drink.
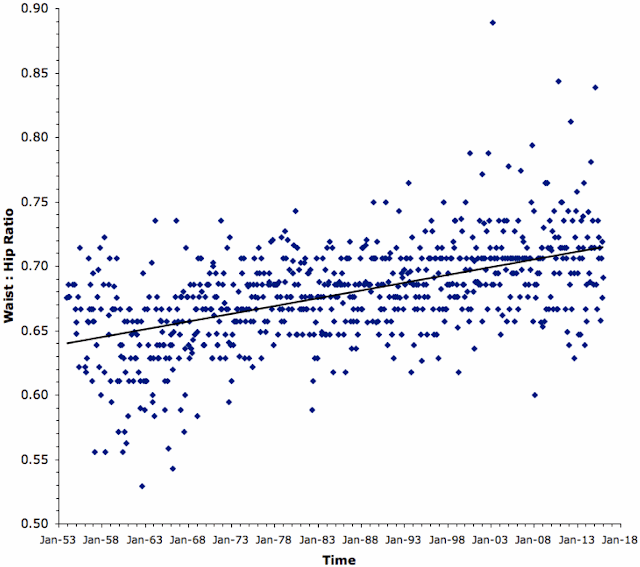
Furthermore, when we plotted Bond's alcohol consumption over time, his intake dropped in the middle of his career but gradually increased towards the end. This consistent but variable lifetime drinking pattern has been reported in patients with alcoholic liver disease.
UK NHS [National Health Service] recommendations for alcohol consumption state that an adult male should drink no more than 21 units a week, with no more than 4 units on any one day, and at least two alcohol free days a week. James Bond's drinking habits are well in excess of each of these three parameters. This level of consumption makes him a category 3 drinker (>60 g alcohol / day) and therefore in the highest risk group for malignancies, depression, hypertension, and cirrhosis. He is also at high risk of suffering from sexual dysfunction, which would considerably affect his womanising.Analyzing the films is more difficult. A number of people have tackled this task, including Nerdist, The Grocer, and Atomic Martinis (now defunct, but repeated on the website of the world's only James Bond Museum, in Sweden), and David Leigh. The basic problem seems to be whether the alcohol is "spotted either in hand, glass or in the background". Also, "The major problem is 007’s frequent enjoyment of multiple bottles of champagne, or portions of bottles of liquor ... it is often impossible to determine exactly how many separate drinks came from a given bottle."
The following infographic (not including the 2015 movie or the unofficial films) is derived from one produced at Buddy Loans. However, some of the people at Reddit were not happy with the original, so it was redesigned, as shown here.
The people at Nerdist took the data from this film infographic, converted it from units of alcohol to grams of alcohol, and then used this to estimate Bond’s total alcohol content. This yields a Blood Alcohol Content of 3.7%. "While some humans have survived a BAC of past 1%, it generally holds that anything past 0.5% will either kill you or leave you seriously poisoned. Therefore ... Bond’s tipsy tally is enough to put a man past a safe limit seven times over."
At The Grocer, they have also pointed out the relative booziness of the various Bond incarnations, by calculating the average intake per film by each actor, in units of alcohol:
| Sean Connery George Lazenby Roger Moore Timothy Dalton Pierce Brosnan Daniel Craig |
11 9 11 4.5 12 20 |
The books are shown in red, and the early films starring Connery and Lazenby are shown in blue (including Connery's later Never Say Never Again). These books and films are almost all at the top and right of the network, indicating that they have a distinct collection of drink types compared to the later films. I suspect that this reflects increasing use of "product placements" in the films. The only book plus movie combination that has similar drinks is You Only Live Twice. Interestingly, the Skyfall movie (from 2012) seems to return to the drinks genre of the earlier works, even though the alcohol consumption is much higher. The most unusual works were the Goldfinger and On Her Majesty's Secret Service books, where a number of drink styles were consumed that appeared nowhere else in the canon.
As noted by Johnson et al. (quoted above):
Despite his alcohol consumption, [Bond] is still described as being able to carry out highly complicated tasks and function at an extraordinarily high level. This is likely to be pure fiction.