Monday, April 29, 2013
A network Tree of Life
In April last year I noted that there are not many images of phylogenetic networks on the internet, suitable for use when an icon or symbol is required, so I provided one (Network road sign). This year I thought that I might point out a far more arty image of "The Tree of Life" that ends up looking more like a network.
The original painting is by Reneé Womack, and prints are available from Fine Art America. You can also have versions with a green or a blue background, but I prefer this one, possibly because it reminds me of the cover of the Tree Thinking book by David Baum and Stacey Smith, which, in turn, is part of the Gustav Klimt mural of the "Tree of Life" (1905-1909).
Wednesday, April 24, 2013
Cloudograms and data-display networks
I have previously noted that splits graphs are a logical way to present the results of Bayesian analyses (We should present bayesian phylogenetic analyses using networks). Bayesian analyses are concerned with estimating a whole probability distribution, rather than producing a single estimate of the maximum probability. Thus, the result of a Bayesian phylogenetic analysis should not be as a single tree (the so-called MAP tree or maximum a posteriori probability tree), but should instead show the probability distribution of all of the sampled trees. This can easily be done with a consensus network, as illustrated by example in the previous blog post.
An interesting alternative way of visualizing the probability distribution of trees is what has been called a Cloudogram, an idea introduced by Remco R. Bouckaert (2010, DensiTree: making sense of sets of phylogenetic trees. Bioinformatics 26: 1372-1373). This diagram superimposes the set of all trees arising from an analysis. Dark areas in such a diagram will be those parts where many of the trees agree on the topology, while lighter areas will indicate disagreement. This idea can be best illustrated by a few published examples.
The first cloudogram is from Figure 4 of Chaves JA, Smith TB (2011) Evolutionary patterns of diversification in the Andean hummingbird genus Adelomyia. Molecular Phylogenetics and Evolution 60: 207-218.
In this case the MAP tree has been superimposed on the cloudogram.
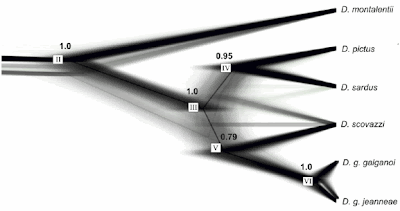
The next one is from Figure 2 of Pabijan M, Crottini A, Reckwell D, Irisarri I, Hauswaldt JS, Vences M (2012) A multigene species tree for Western Mediterranean painted frogs (Discoglossus). Molecular Phylogenetics and Evolution 64: 690-696.
The next one is from Figure 1 of Lerner HR, Meyer M, James HF, Hofreiter M, Fleischer RC (2011) Multilocus resolution of phylogeny and timescale in the extant adaptive radiation of Hawaiian honeycreepers. Current Biology 21: 1838-1844.
In this case the data are more tree-like than the previous two examples.
The final one is from Figure 2 of McCormack JE, Faircloth BC, Crawford NG, Gowaty PA, Brumfield RT, Glenn TC (2012) Ultraconserved elements are novel phylogenomic markers that resolve placental mammal phylogeny when combined with species-tree analysis. Genome Research 22: 746-754.
This analysis involves bootstraps rather than Bayesian samples, showing that the same principle applies.
It would be nice to illustrate this further by direct comparison with a splits graph of the same dataset that I used in the previous blog post. Unfortunately, the computer program available (DensiTree) has the same practical limitation as the SplitsTree program (as mentioned in the previous post) — it does not read the MrBayes ".trprobs" file because it ignores the tree weights. This means that one has to enter the entire treefile (with thousands of trees), and I have not yet done that. Moreover, the program relies very much on having branch lengths for each tree — the output is really quite odd without them, with the taxa appearing in a series of steps rather than connected by straight branches. My previous analysis did not use branch lengths, as they are not needed for the consensus network, in which edge lengths represent support rather than character evolution.
Labels:
Cloudogram,
Posterior probability,
Splits graph
Monday, April 22, 2013
Personal Type I error rates
As usual at the beginning of the week, this blog presents something in a lighter vein. However, this week we depart from phylogenetic networks in the strict sense, and take a humorous look at the broader statistical life of biologists.
Statistics is a curious thing, which allows scientists to make probability errors of two types: Type I (also known as false positives) and Type II (also known as false negatives). Importantly, these errors can accumulate in any one experiment, so that we can also recognize an Experimentwise Error Rate, which is the sum of the individual errors associated with each experimental hypothesis test.
However, what is not widely recognized is that these errors apply in life, as well. In particular, biologists accumulate statistical errors throughout their lives, so that we all have a Personal Lifetime Error Rate.
I once wrote a tongue-in-cheek article about the accumulation of Type I errors throughout the working life of a biological scientist, and the consequences for the experiments conducted by that scientist. This article appeared in 1991 in the Bulletin of the Ecological Society of Australia 21(3): 49–53, which means that I used an ecologist as my specific example of a biologist. The principle applies to all biologists, however.
Since this issue of the Bulletin is not online, presumably no-one has read this article since 1991, although it has recently been referenced on the web (see the sixth comment on this blog post).** You, too, should read it, and so I have linked to a PDF copy [1.7 MB] of the paper:
Personal Type I error rates in the ecological sciences
** Note that I am alternately referred to as an "inveterate mischief maker" and "a very wise man"!
Labels:
computational biology,
inference
Wednesday, April 17, 2013
When is a tree structure a phylogeny?
I have noted before today that many people seem to treat non-biological phylogenetic attributes as being analogous to genotypes whereas most such data are much more similar to phenotypes (eg. False analogies between anthropology and biology; The Music Genome Project is no such thing). This inappropriate analogy can lead to problems, such as incorrect conclusions regarding familial relationships.
In a similar vein, another problem is the appropriation of the word "phylogeny" to refer to non-evolutionary types of tree. A web search for phylogeny will lead you to many sites where the tree structure being referenced is very unlike an evolutionary history.
Systematists have long dealt with this issue as manifest in the confusion between classification and phylogeny. Biological classification is usually treated as most informative (eg. explanatory, predictive) when based on a phylogeny, but a phylogeny is not automatically a classification, and a classification is not automatically a phylogeny.
The best known example is the NCBI Taxonomy, as used by the GenBank database. This is one of the most commonly used classification schemes today, but in bioinformatics it is frequently used as a phylogeny as well as a classification. This is in spite of the fact that NCBI offers the following disclaimer:
The NCBI taxonomy database is not a primary source for taxonomic or phylogenetic information. Furthermore, the database does not follow a single taxonomic treatise but rather attempts to incorporate phylogenetic and taxonomic knowledge from a variety of sources, including the published literature, web-based databases, and the advice of sequence submitters and outside taxonomy experts. Consequently, the NCBI taxonomy database is not a phylogenetic or taxonomic authority and should not be cited as such.The issue here is that the classification is hierarchical and can therefore be expressed as a tree, and the same can said of the nested relationships in a phylogeny. However, not all trees are phylogenies, and the NCBI Taxonomy is a classification that is not necessarily phylogenetic.
More recently, the word phylogeny has been adopted by the computational word to refer to many hierarchical clustering patterns. For example, consider this definition from FreeBase:
The phylogeny pattern is a major pattern within ontology / schema modelling, and is prevalent in many schemas in Freebase. Commonly related are the parent-child pattern and the containment pattern.
In other words, parent-child patterns are phylogenetic, which is literally true as far as it goes, but a two-level hierarchy fits this pattern without being anything more than a trivial phylogeny in the biological sense. An example is the Wikipedia music entries (eg. Rock music), which have a genre and several subgenres, along with fusion genres — this produces a shallow but broad "tree". Indeed, FreeBase has this to say about their own attempt to implement this idea:
One issue is that the some of the data in the music genre hierarchy in Freebase seems to attempt to show a genealogy of genres, rather than family groupings, which is counter to the way that parent and child Media genres are defined.
This seems to be a rather confused set of analogies involving families and genealogies. The false analogy between a tree and a phylogeny seems to have created this confusion. A genealogy expresses family groups (as does a phylogeny), but not all of those potential groups need be expressed in a classification.
It seems to me that it would be simpler for the computational world to refer to a hierarchy rather than a phylogeny.
It seems to me that it would be simpler for the computational world to refer to a hierarchy rather than a phylogeny.
Labels:
Genealogy,
Philosophy,
Phylogeny
Monday, April 15, 2013
A network analysis of Simon and Garfunkel
Every decade or so a record company releases a compilation album from the best-selling musical duo of Paul Frederic Simon and Arthur Ira Garfunkel. There are currently five such albums that have been given a worldwide release:
- Simon and Garfunkel's Greatest Hits (1972)
- The Simon and Garfunkel Collection (1981)
- The Concert in Central Park (1982)
- The Definitive Simon & Garfunkel (1992)
- The Essential Simon & Garfunkel (2003)
This is not bad considering that the duo released only 5 original albums in the first place (Wednesday Morning, 3 A.M.; Sounds of Silence; Parsley, Sage, Rosemary and Thyme; Bookends; Bridge Over Troubled Water), plus one album shared with Dave Grusin (The Graduate). It also means that we are overdue for another compilation.
Each of these compilation albums has been released in a number of different countries, where they have had a greater or lesser success in terms of sales. Some of the relevant information about the resulting chart positions for these 5 albums is available from Wikipedia. This means that we could examine how the different countries compare in their enthusiasm for Simon and Garfunkel's songs.
Unfortunately, not all of the albums were released in all of the countries for which information has been compiled. For example, the USA has data for only 3 of the 5 albums, and Finland and Norway each has data for only 4 of them. Nevertheless, there is complete information for 8 countries, for which I can perform an exploratory data analysis using a network.
As usual, I have used the manhattan distance and a NeighborNet network to produce the graph. The proximity of the countries in the network shows how similarly the records sold — countries near each other had similar sales of the records, while distant countries had different sales patterns.
Only 1 of the 5 albums sold well in all 8 countries: The Concert in Central Park (the chart positions were 1, 1, 1, 2, 3, 5, 5, 6).
Sweden holds a unique position in the network because the populace did not buy The Simon and Garfunkel Collection (chart position 49) but did buy all of the other albums (positions 3-5).
The Netherlands and Japan are linked together in the network because they did not support The Essential Simon & Garfunkel (positions 64 & 104, respectively). Indeed, the Dutch did not like The Definitive Simon & Garfunkel (35), either, while the Japanese seemed to like only The Concert in Central Park (2) and Simon and Garfunkel's Greatest Hits (3).
Australia, Germany and the United Kingdom are linked by sharing their ranking of The Essential Simon & Garfunkel (20-25). France and New Zealand are linked by sharing their ranking of both Simon and Garfunkel's Greatest Hits (16, 22) and The Essential Simon & Garfunkel (33, 38).
Note that it is therefore the sales of The Essential Simon & Garfunkel that has the largest effect on the network pattern:
4 Sweden
20-25 Australia, Germany, UK
33-38 France, New Zealand
64 Netherlands
104 Japan
So, it turns out that the popularity of Simon and Garfunkel does, indeed, have a geographical pattern, although probably not an expected one.
Labels:
EDA,
Simon Garfunkel
Wednesday, April 10, 2013
Highlighting splits in a splits graph
A splits graph is interpreted in terms of splits, or bipartitions, which divide the graph into two non-overlapping parts. If one wishes to refer to particular splits in a graph then one needs a way of highlighting those splits.
This can be done in a number of ways, some of them derived from conventions originating for the presentation of rooted phylogenetic trees. These include highlighting the taxa in one of the partitions, which is analogous to highlighting a clade in a rooted phylogenetic tree. Alternatively, we could colour the edges associated with each of the two partitions, as shown in this previous blog post (How to interpret splits graphs); however, this works only for a single split at a time.
Alternatively, it is also possible to label the edges of the splits themselves, as shown in this previous blog post (Representing evolutionary scenarios using splits graphs). Dabert et al. (Dabert M, Witalinski W, Kazmierski A, Olszanowski Z, Dabert J (2010) Molecular phylogeny of acariform mites (Acari, Arachnida): strong conflict between phylogenetic signal and long-branch attraction artifacts. Molecular Phylogenetics and Evolution 56: 222-241) present another possibility, which is to colour only the edges that separate to two partitions of each split, as shown in the figure.
Labels:
Splits graph
Monday, April 8, 2013
Stick Science Contest
In 2009 and 2010 a group named the Florida Citizens for Science ran what they called a Stick Science Contest, in which the participants contributed stick cartoons that could "be used to educate the general public and especially decision makers about the truth behind one false science argument." In practice, many (but not all) of the contributions concerned mis-interpretations about evolution.
Only the top ten ranked entries for each year were published online. From these finalists, the top three entries received prizes. Here are links to the finalists for 2009 and for 2010.
While the prize-winning entries are good, some of the others are more interesting from the phylogenetic perspective. My favourite one is no. 6 from 2010, by Matthew Bonnan:
I also like Entry F from 2009, by Jan Stephan Lundquist:
This theme was repeated in entry no. 5 from 2010, by Glen Wolfram:
Labels:
Phylogeny
Wednesday, April 3, 2013
Representing evolutionary scenarios using splits graphs
Splits graphs are basically data-display networks, since their intended purpose is to graphically display the patterns of variation in a dataset. These patterns may relate to evolutionary history, or they may not.
A couple of weeks ago I discussed a paper by Myles et al. concerning the genetics of grape cultivars, and this paper provides an interesting example where the patterns of genetic variation seem to be strongly phylogenetic in nature (Myles S, Boyko AR, Owens CL, Brown PJ, Grassi F, Aradhya MK, Prins B, Reynolds A, Chia JM, Ware D, Bustamante CD, Buckler ES. 2011. Genetic structure and domestication history of the grape. Proceedings of the National Academy of Sciences of the USA 108: 3530-3535).
Myles et al. note that: "Archaeological evidence suggests that grape domestication took place in the South Caucasus between the Caspian and Black Seas and that cultivated vinifera then spread south to the western side of the Fertile Crescent, the Jordan Valley, and Egypt by 5,000 y ago." They provide an explicit historical scenario of the evolutionary history of cultivated grapes (Vitis vinifera):
- There are two species involved (V.sylvestris, V.vinifera), both distributed along the eastern and northern part of the Mediterranean basin;
- V.vinifera was domesticated from V.sylvestris in the eastern part of the distribution;
- V.vinifera then spread geographically from east to west;
- This spread was followed by introgression of V.sylvestris into V.vinifera in the western part of their joint distribution.
I constructed a NeighborNet splits graph from these Fst data, as shown in the graph. According to Myles et al., the geographic regions are defined as follows: "east" includes locations east of Istanbul, Turkey; "west" includes locations west of Slovenia, including Austria; and "central" refers to locations between them.
Note, however, that one cannot infer the scenario from the splits graph, because the data analysis is not intended for direct evolutionary inference. The graph is undirected, and there are therefore several possible scenarios that could be derived from the graph. For example, the graph shown is also compatible with the domestication of V.vinifera from V.sylvestris in the western part of the distribution.
Thus, a splits graph can be used to suggest scenarios (ie. hypothesis generation) and it can be used to test scenarios (hypothesis testing), but the latter is a weak test because there will always be several phylogenetic scenarios with which it is compatible.
Labels:
Splits graph
Monday, April 1, 2013
Empedocles, Lucretius and lateral gene transfer
Empedocles (c. 490–430 BCE) and Lucretius (c. 99-55 BCE) have been credited with first articulating the theory of "survival of the fittest" (Sedgley 2003). However, this is of interest only to Darwinian scholars, who focus solely on trees. What is of more interest to scholars of phylogenetic networks is that these same two philosophers have also been credited with first suggesting the doctrine of horizontal gene transfer (Wilkins 2009). Gene transfer is, of course, an important source of reticulate evolution.
Empedocles was a Greek philosopher, a citizen of what is now Agrigento, in Sicily. He is perhaps most famous for first outlining the elemental theory of the physical world (ie. Air, Earth, Fire, Water). Moreover, he identified two fundamental forces, which he called love and strife. Love is the force that brings objects together, while Strife is the force that drives them apart. Empedocles postulated that the universe was once condensed into a tight sphere by the force of love, and strife later exploded this into an expanding mass. This has been seen as a forerunner of modern ideas about the Big Bang and the subsequent expanding universe.
More importantly for our purposes, Empedocles had a physical theory about the random development of living forms. According to this theory, Life first emerged as a collection of disassociated body parts, which wandered about on their own, without the intervention of divine power. These were not parts severed from previously complex beings, but each functioned in its own right as an independent "single-limbed" being. Complex creatures were then created by the accidental combination of these disparate limbs and organs. If the correct parts combined, then the creature would survive and go on to found a species, but if the wrong combination occurred then the creature would perish — only those with the most suitable combinations survived, by a process that we now call natural selection.
Empedocles' hypothesized hybrid creatures were literally mocked by later Greek philosophers, notably Aristoteles (384-322 BCE) and Epicurus (341-270 BCE), and their followers. They derided these monsters as "roll-walking creatures with hands not properly articulated or distinguishable" and as "ox-headed man-creatures". It was Lucretius who resurrected Empedocles' idea, in the fifth part of his only known work (the poem De Rerum Natura), which was about the beliefs of Epicureanism — Lucretius was the first writer to introduce Roman readers to Epicurean philosophy.
Titus Lucretius Carus was a Roman poet and philosopher, apparently resident in Rome itself. He is perhaps most famous for his atomistic view of the physical world (everything is built up from collections of indivisible particles). More importantly for our purposes, Lucretius expounded a similar theory to that of Empedocles, namely that originally a set of randomly composed monsters sprang up, of which only the fittest survived. However, whereas Empedocles described isolated limbs as the starting point, Lucretius described whole organisms with defective combinations of body parts (what we would now call congenital defects), so that his maladapted creatures were formed at the atomic level rather than at the macroscopic level of whole limbs. Also, in Lucretius' theory there was apparently no inter-species mingling of limbs, as there was in Empedocles' version.
These two related theories of zoogony appear to have lain dormant for a couple of thousand years, crushed under the iron fist of both Aristoteleanism and the early Christian era. Even into the 1900s, biology could be best described as being essentially an extension of Aristoteles' philosophical ideas (Mayr 1982). Nevertheless, slowly the idea of natural selection was re-introduced to biology, notably with the work of Étienne Geoffroy Saint-Hilaire (1772-1844), and culminating in the work Alfred Russel Wallace (1823-1913) and Charles Robert Darwin (1809-1882).
However, even after the introduction of this evolutionary idea, the focus was on the inheritance of morphological modifications, not on the admixture of parts inherited from different organisms; and so only half of Empedocles' ideas were accepted.
It took until the dawn of the 20th century for the Russian lichenologist Constantin Sergeevich Mereschkowsky (1855-1921) to first outline a cellular version of Empedocles' vision. It had recently been shown that lichens involve a symbiotic relationship between fungi and algae, very much along the lines first envisioned more than 2,200 years before. Mereschkowsky extended this idea to the sub-cellular level, with the explicit goal of explaining the evolutionary development of land plants from algae-like forms of life, postulating that chloroplasts originated as symbiotic blue-green algae. The German histologist Richard Altman (1852-1900) had already hinted that what we now call mitochondria (he called them bioblasts) are bacterial symbionts. It was some time later that the American anatomist Ivan Emanuel Wallin (1883-1969) published Symbionticism and the Origin of Species, in which he explicitly suggested that symbiotic bacteria have played a fundamental role in the evolution of species.
This development culminated in the suggestion that genes themselves can be transferred between distant organisms, thus bringing thought down to the atomistic level envisioned by Lucretius. This revealed the hybrid nature of many genomes, even in situations where phenotypic admixture is not manifest. The first description of horizontal gene transfer is usually credited to Victor J. Freeman (in 1951), who demonstrated that the transfer of a viral gene into a bacterium could create a virulent strain from a non-virulent strain. Since then, lateral gene transfer has been widely reported as an important component of prokaryote evolution; and it has increasingly been reported in eukaryotes as well.
We have thus come full circle. Empedocles first introduced the theory of "survival of the fittest", which took nearly 2,300 years to be re-discovered by science, as well as outlining the basic concept of "horizontal gene transfer", which took an extra century for its renaissance.
All of the information presented here is factually correct. However, only on All Fool's Day can the facts be combined in this outrageous way, and such a history be told with a straight face.
References
Mayr E. (1982) The Growth of Biological Thought: Diversity, Evolution and Inheritance. Belknap Press, Cambridge MA.
Sedgley D. (2003) Lucretius and the new Empedocles. Leeds International Classical Studies 2.4.
Wilkins J.S. (2009) New work on lateral transfer shows that Darwin was wrong. ScienceBlogs Evolving Thoughts March 31 2009.
Labels:
HGT network,
History
Subscribe to:
Posts (Atom)