Wednesday, October 29, 2014
Uncertainty in multiple sequence alignments
It is well known that reticulations in phylogenetic networks can reflect variation in data sets from many sources, not only gene flow during evolutionary history. These other sources are presumably unwanted in the analysis when they are due to estimation errors. Such errors include incorrect data, inappropriate sampling, and model mis-specification.
For molecular data, one of the more obvious sources of model mis-specification is an incorrect multiple sequence alignment. This reflects wrong assessments of primary homology among the characters, so that the wrong residues are aligned in the columns. This particular issue seems not to have been addressed in the network literature in any systematic way.
However, it is obviously rather important. After all, who needs a phylogenetic network that reflects mis-alignment rather than evolutionary history? One approach to this issue would be to have some sort of measurement of our confidence in the alignment columns, which could be taken into account when the network is constructed.
One practical problem with this approach is that there has been a veritable cottage industry developing such measurements, which would need to be assessed for their suitability. So, I thought that I might list some of them here, along with a brief description of what they measure. The list is comprehensive but not necessarily exhaustive — it consists of ones for which there was at some stage a computer program (there are others that have never been named). Most of the methods are designed specifically for amino-acid sequences, so that not all of them can be used for nucleotides.
There are basically two types of measurement: (1) quantitative scoring schemes, which provide a reliability score for each aligned position, and (2) selection schemes, which select a subset of the aligned positions as being reliably aligned. So, I have divided the list roughly into these two groups.
QUANTITATIVE
WET
Dopazo J (1997) A new index to find regions showing an unexpected variability or conservation in sequence alignments. Computer Applications in the Biosciences 13: 313-317.
— evolutionary index is based on conservativeness of amino acid differences as predicted from nucleotide differences
ClustalX
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL-X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research 25: 4876-4882.
— quality is based on conservativeness of amino acid differences
Coffee
Notredame C, Holm L, Higgins DG (1998) COFFEE: an objective function for multiple sequence alignments. Bioinformatics 14: 407-422.
— score represents consistency among global and local alignments
AL2CO
Pei J, Grishin NV (2001) AL2CO: calculation of positional conservation in a protein sequence alignment. Bioinformatics 17: 700-712.
— conservation is based on weighted entropy
BAli-Phy
Redelings BD, Suchard MA (2005) Joint Bayesian estimation of alignment and phylogeny. Systematic Biology 54: 401-418.
— approximate probability that the letter is homologous to the ancestral residue in its column
MUMSA
Lassmann T, Sonnhammer EL (2005) Automatic assessment of alignment quality. Nucleic Acids Research 33: 7120-7128.
— consistency based on overlap of alignments from several programs
HoT score
Landan G, Graur D (2007) Heads or tails: a simple reliability check for multiple sequence alignments. Molecular Biology and Evolution 24: 1380-1383.
— measures uncertainty due to co-optimal alignments
FSA
Bradley RK, Roberts A, Smoot M, Juvekar S, Do J, Dewey C, Holmes I, Pachter L (2009) Fast Statistical Alignment. PLoS Computational Biology 5: e1000392.
— several scores based on HMM consistency, certainty, expected accuracy, expected sensitivity, expected specificity
Guidance
Penn O, Privman E, Landan G, Graur D, Pupko T (2010) An alignment confidence score capturing robustness to guide tree uncertainty. Molecular Biology and Evolution 27: 1759-1767.
— robustness to guide tree uncertainty
PSAR
Kim J, Ma J (2011) PSAR: measuring multiple sequence alignment reliability by probabilistic sampling. Nucleic Acids Research 39: 6359-6368.
— agreement with probabilistic sampling of suboptimal alignments
Zorro
Wu M, Chatterji S, Eisen JA (2012) Accounting for alignment uncertainty in phylogenomics. PLoS One 7: e30288.
— pair Hidden Markov Model to model the sequence evolution and uses the model to calculate the posterior probabilities that residues of a column are correctly aligned
TCS
Chang J-M, Di Tommaso P, Notredame C (2014) TCS: a new multiple sequence alignment reliability measure to estimate alignment accuracy and improve phylogenetic tree reconstruction. Molecular Biology and Evolution 31: 1625-1637.
— transitive consistency score is an extended version of the Coffee scoring scheme
SELECTION
OSA
Martin MJ, Gonzâlez-Candelas F, Sobrino F, Dopazo J (1995) A method for determining the position and size of optimal sequence regions for phylogenetic analysis. Journal of Molecular Evolution 41: 1128-1138.
— locates the smallest blocks with similar pairwise genetic distances to the whole alignment
Gblocks
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Molecular Biology and Evolution 17: 540-552.
— selected blocks are based on conservation of identity
SOAP
Löytynoja A, Milinkovitch MC (2001) SOAP, cleaning multiple alignments from unstable blocks. Bioinformatics 17: 573-574.
— stability is measured with respect to variation in the Clustal gap-opening and gap-extension penalties
NorMD
Thompson JD, Plewniak F, Ripp R, Thierry J-C, Poch O (2001) Towards a reliable objective function for multiple sequence alignments. Journal of Molecular Biology 314: 937-951.
— normalized mean distance is based on pairwise distances
Shift score
Cline M, Hughey R, Karplus K (2002) Predicting reliable regions in protein sequence alignments. Bioinformatics 18: 306-314.
— uses information from near-optimal alignments
LumberJack
Lawrence CJ, Zmasek CM, Dawe RK, Malmberg RL (2004) LumberJack: a heuristic tool for sequence alignment exploration and phylogenetic inference. Bioinformatics 20: 1977–1979.
— identifies blocks that have their phylogenetic tree being most similar to that of the whole alignment
Noisy
Dress AW, Flamm C, Fritzsch G, Grünewald S, Kruspe M, Prohaska SJ, Stadler PF. (2008) Noisy: identification of problematic columns in multiple sequence alignments. Algorithms in Molecular Biology 3: 7.
— identification of phylogenetically uninformative homoplastic sites from compatibilities in a circular split system
trimAl
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25: 1972-1973.
— proportion of sequences with a gap, level of amino acid similarity, level of consistency across different (user-provided) alignments
SVM
Blouin C, Perry S, Lavell A, Susko E, Roger AJ. (2009) Reproducing the manual annotation of multiple sequence alignments using a SVM classifier. Bioinformatics 25: 3093-3098.
— support vector machine reproduces manual annotations from other alignments
BMGE
Criscuolo A, Gribaldo S (2010) BMGE (Block Mapping and Gathering with Entropy): a new software for selection of phylogenetic informative regions from multiple sequence alignments. BMC Evolutionary Biology 10: 210.
— calculates entropy-like scores weighted by similarity matrices
Aliscore
Kück P, Meusemann K, Dambach J, Thormann B, von Reumont BM, Wägele JW, Misof B (2010) Parametric and non-parametric masking of randomness in sequence alignments can be improved and leads to better resolved trees. Frontiers in Zoology 7: 10.
— consensus profiles identify dominating patterns of nonrandom similarity
SR
Rajan V (2013) A method of alignment masking for refining the phylogenetic signal of multiple sequence alignments. Molecular Biology and Evolution 30: 689-712.
— compatible subsplits define clusters of sites which are then removed based on evolutionary rate
Labels:
Alignment
Monday, October 27, 2014
Predecessors of Charles Darwin
Charles Darwin and Alfred Russel Wallace are usually credited with independently developing the idea that natural selection could be the important process by which new species arise, although history has apportioned most of the fame to Darwin alone.
In the first edition of his most famous book Darwin (1859) cited no sources, and credited no-one except Thomas Malthus as a source of ideas. He was criticized for this, and from the third edition onwards he provided a historical essay mentioning a few more names.
The basic issue is that the idea of natural selection had been "in the air" for more than half a century, but only with respect to within-species variation. It was Darwin and Wallace who took the leap to consider between-species variation, on the basis that there is no historical boundary defining species — all individuals trace their ancestry back through a whole series of ancestors, including those who existed before the origin of their current species. That is, phylogenies trace back to the origin of life not just to the origin of each species.
So, who were the people who published, however briefly, a comment noting the idea of within-species natural selection? Joachim Dagg, of the Natural History Apostils blog, has recently been writing a series of posts discussing many of those publications that contain a clear description of selection. Here I have provided a convenient overview, in time order, with links to Joachim's blog for those of you who want more information.
Joseph Townsend
- (1786, republished in 1817) A Dissertation on the Poor Laws, by a Well-wisher to Mankind. London: Ridgways.
Link 1 - Link 2
James Hutton
- (1794) Investigation of the Principles of Knowledge and of the Progress of Reason, from Sense to Science and Philosophy. Volume 2. Edinburgh: Strahan & Cadell. [section 13, chapter 3]
Link 1
William Charles Wells
- (1813) An Account of a White Female, Part of Whose Skin Resembles that of a Negro. [talk]
- (1818) Two Essays: One Upon Single Vision with Two Eyes; the other on Dew. [plus] An Account of a Female of the White Race of Mankind, Part of Whose Skin Resembles that of a Negro. Edinburgh: Archibald Constable.
Link 1 - Link 2
Joseph Adams
- (1814) A Treatise on the Supposed Hereditary Properties of Diseases. London: J. Callow.
Link 1 - Link 2 - Link 3
Patrick Matthew
- (1831) On Naval Timber and Arboriculture; with Critical Notes on Authors who have Recently Treated the Subject of Planting. Edinburgh: Adam Black.
Link 1 - Link 2 - Link 3 - Link 4 - Link 5
You can learn more about him at The Patrick Matthew Project.
John C. Loudon
- (1832) [Book review of] Matthew, Patrick: On Naval Timber and Arboriculture; with Critical Notes on Authors who have recently treated the Subject of Planting. The Gardener's Magazine 8: 702-703.
Link 1 - Link 2
Edward Blyth
- (1835) An attempt to classify the "varieties" of animals, with observations on the marked seasonal and other changes which naturally take place in various British species, and which do not constitute varieties. The Magazine of Natural History 8: 40-53.*
- (1836) Observations on the various seasonal and other external changes which regularly take place in birds, more particularly in those which occur in Britain; with remarks on their great importance in indicating the true affinities of species; and upon the natural system of arrangement. The Magazine of Natural History 9: 393-409.*
- (1837) On the psychological distinctions between man and all other animals; and the consequent diversity of human influence over the inferior ranks of creation, from any mutual or reciprocal influence exercised among the latter. The Magazine of Natural History, new series, 1: 1-9.*
Link 1
Herbert Spencer
- (1852) A theory of population, deduced from the general law of animal fertility. Westminster Review 57: 468-501.
Link 1
* Full title: The Magazine of Natural History and Journal of Zoology, Botany, Mineralogy, Geology, and Meteorology
Wednesday, October 22, 2014
Is phylogenomics tree-like?
Phylogenomics, the idea of applying genomic data to phylogenetic studies, has been around for quite a while now (Eisen 1998), although it was probably Rokas et al. (2003) who drew the first widespread attention among phylogeneticists. Molecular phylogenetics started off using the sequence of a single locus (often small-subunit rRNA) as the data, and slowly progressed from there to multiple loci. Currently, it is considered good practice to use half-a-dozen loci, sampling the main genomes (nucleus, mitochondrion, plastid); and genomics offers the possibility of a fast and cost-effective means of generating large amounts of multi-locus sequence data.
Review papers are beginning to appear based explicitly on next-generation sequencing (NGS), such as those of Lemmon & Lemmon (2013) and McCormack et al. (2013), replacing the earlier work of Philippe et al. (2005), and there are suggestions for how phylogenetics analyses might need to change in response to NGS data (Chan and Ragan 2013). These all treat phylogenomics as being very similar to traditional molecular phylogenetics, in the sense that many people are expecting phylogenomics to provide tree-like resolution of questions that remain unresolved with the current smaller datasets. In the words of Rokas et al. (2003), phylogenomics is intent on "resolving incongruence in molecular phylogenies". That is, incongruent gene trees are seen as the major obstacle to be overcome by phylogenetics data analysis (see also Jeffroy et al. 2006).
However, this might be a naive expectation. After all, the existing phylogenetic conflicts are there for a reason. If we cannot resolve certain parts of organismal history in terms of a phylogenetic tree when we use the current levels of multi-locus data (say <10 loci), then there is no real reason to think that this will happen just because we increase the number of loci. There are plenty of other reason for incongruence among genes, the most obvious one being that the history is not tree-like in the first place. The advantage of phylogenomics, then, would be its ability to clarify the phylogenetic history rather than to resolve incongruence.
There are now quite a few published empirical phylogenomic studies, which allows us to provide a preliminary answer to the question about whether phylogenomic patterns are tree-like or not. There are a few published studies where the authors claim resolution in terms of a tree, as least for part of their phylogeny (e.g. Wang et al. 2012), but it seems to me that there are far more studies where the incongruence remains even with genomic data. Below, I briefly introduce a few arbitrarily chosen examples.
So, complex genealogical problems often remain complex even after using genomic data. We haven’t "solved" any of the so-called genealogy problems, we have simply made clear in what way they are complex. That is, genomics data generally reveal reticulate evolutionary histories, not simple tree-like ones.
This leads me to conclude that phylogenomics is about reticulate evolution, and it is thus time for phylogeneticists to abandon trees as a model for genealogies. We have probably already resolved most of the simple tree-like genealogical patterns, using non-genomic data, and from here on we will be using genomics to study gene flow in addition to parental gene inheritance.
Examples
(1) Galtier and Daubin (2008) were among the earliest researchers to try to "deal with incongruence in phylogenomic analyses", and one of their examples was the long-standing problem deciphering the relationships among the closest relatives of humans. However, the genomics data make it clear that, while humans share slightly more genes with chimpanzees than with other great apes, we still share some with gorillas but not chimpanzees, and with orangutans but not either chimpanzees or gorillas. Also, chimpanzees share some genes with gorillas that we do not share. The situation is now clearer, but the tree incongruence remains.
(2) At the same time, Kuo et al. (2008) looked at the then-available genomes for members of the Apicomplexa, which are unicellular eukaryotic parasites. The genomic data confirmed the current groupings of Haemosporidians, Piroplasmids and Coccidians (shown as branches with high support in the diagram) but completely failed to resolve the relationships between these groups (shown as branches with low support). Things are no better today, when we have at least some data for 35 genomes.
(3) The relationships among mammal superorders, particularly the placentals, has been a ongoing area of debate. I have already covered this in some previous blog posts, notably Conflicting placental roots: network or tree? and Why are there conflicting placental roots? There are three possible ways of resolving a tree at the root of the placental phylogeny, and genomic datasets seem to support all three of them — the published different trees are therefore based on variation in the model used for data analysis. As Hallström and Janke (2010) have noted, there was probably incomplete lineage sorting and hybridization in the early placental mammalian divergences, rather than a truly tree-like history.
(4) Dell'Ampio et al. (2014) looked at the phylogenetic relationships of the wingless insects, and tried to come to grips with the incongruence among genes. They considered three main tree-based hypotheses for the relationships, and found that genomic support was pretty evenly spread among the three topologies. They dryly note that after their hard work the relationships "are still considered unresolved."
(5) Relationships among hominids have been a popular study for many years, and not unexpectedly there has been a burst as a result of genomic data, especially as there are now SNP micro-arrays available to simplify the data collection. I have covered this in previous posts, as well, notably Why do we still use trees for the Neandertal genealogy? The bottom line is that the genomic data provide evidence of extensive introgression (or admixture) between humans and their nearest relatives throughout their time of co-existence. This example is from Reich et al. (2011).
References
Chan CX, Ragan MA (2013) Next-generation phylogenomics. Biology Direct 8: 3.
Dell'Ampio E, Meusemann K, Szucsich NU, Peters RS, Meyer B, Borner J, Petersen M, Aberer AJ, Stamatakis A, Walzl MG, Minh BQ, von Haeseler A, Ebersberger I, Pass G, Misof B (2014) Decisive data sets in phylogenomics: lessons from studies on the phylogenetic relationships of primarily wingless insects. Molecular Biology and Evolution 31: 239-249.
Eisen JA (1998) Phylogenomics: improving functional predictions for uncharacterized genes by evolutionary analysis. Genome Research 8: 163-167.
Galtier N, Daubin V (2008) Dealing with incongruence in phylogenomic analyses. Philosophical Transactions of the Royal Society of London, Series B: Biological Sciences 363: 4023-4029.
Hallström BM, Janke A (2010) Mammalian evolution may not be strictly bifurcating. Molecular Biology and Evolution 27: 2804-2816.
Jeffroy O, Brinkmann H, Delsuc F, Philippe H (2006) Phylogenomics: the beginning of incongruence? Trends in Genetics 22: 225-231.
Kuo C-H, Wares JP, Kissinger JC (2008) The Apicomplexan whole-genome phylogeny: an analysis of incongruence among gene trees. Molecular Biology and Evolution 25: 2689-2698.
Lemmon EM, Lemmon AR (2013) High-throughput genomic data in systematics and phylogenetics. Annual Review of Ecology, Evolution, and Systematics 44: 99-121.
McCormack JE, Hird SM, Zellmer AJ, Carstens BC, Brumfield RT (2013) Applications of next-generation sequencing to phylogeography and phylogenetics. Molecular Phylogenetics and Evolution 66: 526-538.
Philippe H, Delsuc F, Brinkmann H. Lartillot N (2005) Phylogenomics. Annual Review of Ecology, Evolution, and Systematics 36: 541-562.
Reich D, Patterson N, Kircher M, Delfin F, Nandineni MR, Pugach I, Ko AM, Ko Y-C, Jinam TA, Phipps ME, Saitou N, Wollstein A, Kayser M, Pääbo S, Stoneking M (2011) Denisova admixture and the first modern human dispersals into Southeast Asia and Oceania. American Journal of Human Genetics 89: 516-528.
Rokas A, Williams BL, King N, Carroll SB (2003) Genome-scale approaches to resolving incongruence in molecular phylogenies. Nature 425: 798-804.
Wang N, Braun EL, Kimball RT (2012) Testing hypotheses about the sister group of the Passeriformes using an independent 30 locus dataset. Molecular Biology and Evolution 29: 737-750.
Monday, October 20, 2014
Beer family trees
Some time ago I wrote a blog post about The bourbon family forest, which contained a collection of trees that, rather than being genealogical trees, instead showed the corporate ownership of American whiskey.
Here is a similar arrangement for "the six companies that make 50% of the world's beer", produced by David Yanofsky at the Quartz blog. As before, the vertical axis is actually a time scale, but the trees are only marginally family trees in the genealogical sense. Note that there is a reticulation between two of the trees for the "Scottish & Newcastle" entry, although this was apparently followed immediately by a subsequent divergence.
Nevertheless, roughly the same sort of information could actually be presented as proper genealogies. Here is an example form Philip Howard's blog, restricted to American beer. Note that the genealogies refer to the joining of branches through time, rather than their splitting. There are two reticulation events, one of which also refers to the "Scottish & Newcastle" entry.
It is also worth noting the use of other types of network by Philip Howard, to look at:
- Visualizing ownership in the soft drink industry
- Concentration in the U.S. beer industry
- Concentration in the U.S. wine industry
Labels:
Beer
Wednesday, October 15, 2014
Open problems in phylogenetics
Periodically, mathematicians and other computationalists produce lists of what they refer to as "Open Problems" in their particular field. Phylogenetics is no exception. We have had a few on this blog before today (e.g. An open question about computational complexity; Phylogenetic network Millennium problems).
I thought that I should draw your attention to the fact that last year, Barbara Holland produced a few of her own (2013. The rise of statistical phylogenetics. Australian and New Zealand Journal of Statistics 55: 205-220). These are:
Open problem 1: What is the natural analogue of a confidence interval for a phylogenetic tree?
Open problem 2: What are useful residual diagnostics for phylogenetic models?
Open problem 3: What makes a good phylogenetic model?
Open problem 4: Should DAGs be acceptable objects for inference or should network methods be restricted to exploratory data analysis?
It is obviously the latter problem that is of most interest to us here:
DAGs [directed acyclic graphs] can be constructed by beginning with a good tree and then progressively adding edges until the fit between the model and the data is deemed good enough or there is no sufficient improvement in fit by continuing to add edges. The trouble with using DAGs to define mixture models is that this approach doesn’t actually capture the biological processes of interest within the model. The sorts of things we’d like the data to tell us are what is the relative rate of recombination events or hybridisation events to mutation events or speciation events. The danger with using phylogenetic networks in an "add an extra edge until the fit is good enough" approach is that by giving ourselves the capacity to explain everything we risk explaining nothing. At some point have we stopped doing inference and got back to just summarising our data?
In phylogenetics we rely on our models for their explanatory power — in the context of network evolution we need to make careful decisions about what biological processes should be included within the model such that inferences about reticulate (non-treelike) processes of evolution can be brought within the realm of stochastic uncertainty rather than being left as a source of inductive uncertainty. This is not a straightforward task, and will require the collaboration of evolutionary biologists and statisticians.One of the principal issues here is that it is almost impossible to consistently distinguish one reticulation process from another based on the structure of the resulting network. These processes all produce gene flow in the biological world, and they all appear as reticulations in the graphical representation of a network. In practice, phylogenetic analysis may boil down to only two biological processes in the model (vertical gene inheritance and horizontal gene flow), followed by biologists trying to sort out the details with post hoc analyses. Deep coalescence and gene duplication are part of the vertical inheritance, while hybridization, introgression, horizontal gene flow and recombination are part of gene flow. It would be nice to think that this model would simplify network analyses.
Labels:
open problems
Monday, October 13, 2014
The phylogeny of plastic bag ties
Some years ago Larisa Lehmer, Bruce Ragsdale, John Daniel, Edwin Hayashi and Robert Kvalstad published a medical report about an ingested plastic bag closure caught in someone's colon (Plastic bag clip discovered in partial colectomy accompanying proposal for phylogenic plastic bag clip classification. BMJ Case Reports 2011). This sounds quite painful.
What is more interesting, though, is that the report was accompanied by a phylogenetic and taxonomic evaluation of plastic ties in general, which the authors named Occlupanids.
Note that the proposed morphological changes in the phylogeny match Cope's Rule of phyletic size increase, as discussed in a previous blog post (Steven Jay Gould was wrong).
Shortly afterwards, one of the authors, John Daniel, set up a web page with a more detailed analysis, under the guise of the Holotypic Occlupanid Research Group (HORG).
Among a lot of other interesting information, there is a revised phylogenetic analysis.
Given the data, it seems fairly clear that the genealogical relationship among these objects is reticulate, and that the trees should thus actually be networks. This follows from the simple fact that these phylogenies are rather uninformative (they are bushes showing a few character transformation series). Also, note that contemporary taxa are ancestors, so that the diagrams are more like population networks than species networks.
These ties are used for packets of sliced bread (a relatively recent invention), and so there has been an explosion of Occlupanid forms as they occupy a new adaptive zone. This is a classic instance of recent speciation that is not yet complete. Occlupanids have now reached pest proportions, except where governments have instituted erradication programmes (such as Europe, where they are no longer found).
Part of the difficulty of analysis is that the objects shown constitute only a small part of the known diversity of Occlupanids (e.g. see this photo and this one). There are a number of manufacturers, and their products constitute separate historical lineages. Morphological features have been transferred from one lineage to another, which is a classic case of reticulate history that has not been taken into account in the above phylogenies.
Indeed, the HORG page is not the only detailed web resource about bread ties — see also the now-defunct but fascinating Transactoid page.
Labels:
Bread ties,
Phylogeny,
Steven Jay Gould
Wednesday, October 8, 2014
Thoroughbred horses and reticulate pedigrees
I noted recently that the best documented human genealogies are those for the various Anabaptist populations (including the Mennonites, Hutterites and Amish) (The importance of the Amish for reticulate genealogies). They have mostly closed populations (ie. marriages occur solely within a population), and they are thus inbred, and most importantly they maintain detailed written genealogies. This makes them ideal for genealogical studies involving reticulation, including being a source of "known" reticulate histories for testing network algorithms.
If we move outside of Homo sapiens then a genealogy that is equally well documented (if not better) is that of English Thoroughbred horses. This breed was developed as a result of the enthusiasm of the British aristocracy for racing in the 17th century. Thoroughbred pedigree records are regarded as the most comprehensive records detailing ancestral relationships among domestic animal breeds, and they have been formally catalogued since the appearance of the first edition of the General Stud Book in 1791.
As noted by Binns et al. (2011):
The Thoroughbred horse breed was established in England in the early 1700s based on crosses between stallions of Arabian origin and indigenous mares. The founder population was small, with all current males tracing back to one of three stallions, the Godolphin Arabian, the Byerley Turk and the Darley Arabian; in contrast, on the female side, about 70 foundation mares have been identified. A stud book for Thoroughbred horses was initiated in 1791, and pedigree records for the breed, which now number about five hundred thousand horses, are maintained by Thoroughbred registries worldwide.For the males, the story is continued by Bower et al. (2012):
All living Thoroughbreds trace paternally to just three stallions imported into England in the late 17th and early 18th centuries: Byerley Turk (1680s), Darley Arabian (1704) and Godolphin Arabian (1729). Furthermore, a small number of stallions exerted disproportionate influence on early Classic races resulting in their greater popularity at stud. Therefore, the Thoroughbred gene pool has been restricted by small foundation stock and subsequent limited paternal contributions as a result of sire preference and selection. [Our] historic samples were related largely via the Darley Arabian sire line to which 95% of all living Thoroughbreds can be traced in their paternal lineage.Actually, 95% of living Thoroughbreds trace their male lineage to Eclipse (1764), a great-great grandson of the Darley Arabian, so that it is Eclipse who appears as the progenitor in most published genealogies (eg. see the one below). Information about these early males is available at this Thoroughbred Heritage page.
Females have been of less interest to horse breeders, and so in many cases we do not know who they were, and in many others we have only a generic name (eg. "Miss Darcy's pet mare", "old Montagu mare", "royal mare", etc). This means that in modern horses there is a high level of mtDNA diversity due to multiple female lineages but there is very little sequence diversity on the Y chromosome (Wallner et al. 2013). Nevertheless, Hill et al. (2002) have tried to trace the influence of the early females on current genotypes, singling out 19 of them as having large influence (on the mitochondrial genealogy), while Bower et al. (2011) provide a broader analysis. Information about these early females is available at this Thoroughbred Heritage page.
The relevance of this information for genealogy studies is that it tells us the Thoroughbred genealogy is effectively closed (little outside breeding), and it is thoroughly documented. This is potentially another source of known reticulate genealogies.
Of particular interest to horse breeders is inbreeding (see Binns et al. 2012). In suitable doses this is seen as a Good Thing, because it can produce the homozygous appearance of desirable racing characteristics. However, inbreeding should not be too recent. For example, if we look at the list of the Blood-Horse Top 100 Thoroughbreds of the 20th Century then none of them have inbreeding in the previous generation and only one has inbreeding in the one before that. However, 54% of the horses have inbreeding in the fourth ancestral generation, and 18% in each of the third and fifth generations. Only 9 horses had no inbreeding during the five previous generations.
For this reason, the standard version of horse genealogies only goes back five generations. This is the stage at which the inbreeding coefficient becomes <1% — inbreeding earlier than five generations has no practical effect on homozygosity. There are potentially 32 ancestors in the 5th generation, contributing 1/32=3% of the DNA on average. This inbreeding is of interest to us because it creates extensive reticulation in horse genealogies.
Pedigree data are readily available at sites like Pedigree Online. Pedigrees are usually drawn as treemaps (see the blog post Trees, treemaps and networks) with horses being repeated as often as necessary to be able to draw the network as a tree (see the blog post Reducing networks to trees). Here is a typical example, for the horse Maddox, without recent inbreeding. Males are in blue and females pink, with the parents at the left and their ancestors proceeding to the right.
Here is an example, for the horse Induna Mkubwa, with inbreeding in the 3rd+4th ancestral generations (highlighted in purple) and also in the 4th+5th generations (in green). Note that the horse Be My Chief is also inbred, in his 4th ancestral generation (in green).
Clearly, this second genealogy should more properly be drawn as a reticulating network. Once this sort of thing is done the reticulations become obvious. Here is an example network for the horse known as Roberto. The horses are numbered in the manner conventional for human pedigrees, with the males on the left of each pair. This is about as complex as it gets for these horses; and you will note that there are only two-thirds of the "expected" number of ancestors.
Finally, here is an example network from the paper by Bower et al. (2012), covering a longer time period but restricted to selected male horses (ie. the female lineages that lead to the reticulation are not named).
Thanks to Induna Mkubwa for the idea for this post.
References
Binns MM, Boehler DA, Bailey E, Lear TL, Cardwell JM, Lambert DH (2012) Inbreeding in the Thoroughbred horse. Animal Genetics 43: 340-342.
Bower MA, Campana MG, Whitten M, Edwards CJ, Jones H, Barrett E, Cassidy R, Nisbet RE, Hill EW, Howe CJ, Binns M. (2011) The cosmopolitan maternal heritage of the Thoroughbred racehorse breed shows a significant contribution from British and Irish native mares. Biology Letters 7: 316-320.
Bower MA, McGivney BA, Campana MG, Gu J, Andersson LS, Barrett E, Davis CR, Mikko S, Stock F, Voronkova V, Bradley DG, Fahey AG, Lindgren G, MacHugh DE, Sulimova G, Hill EW (2012) The genetic origin and history of speed in the Thoroughbred racehorse. Nature Communications 3: 643.
Hill EW, Bradley DG, Al-Barody M, Ertugrul O, Splan RK, Zakharov I, Cunningham EP (2002) History and integrity of thoroughbred dam lines revealed in equine mtDNA variation. Animal Genetics 33: 287-294.
Wallner B, Vogl C, Shukla P, Burgstaller JP, Druml T, Brem G (2013) Identification of genetic variation on the horse Y chromosome and the tracing of male founder lineages in modern breeds. PLoS One 8: e60015.
Monday, October 6, 2014
Network map of the Ukraine
There is a tolerably well-known exercise for illustrating the graphical superiority of a Non-Metric Multidimensional Scaling (NMDS) ordination over a Principal Components Analysis (PCS) ordination. The latter is often subject to distortions, so that the relative positions in the scatter-plot of points do not represent the original measured distances between those points (see the post Distortions and artifacts in Principal Components Analysis analysis of genome data). The exercise consists of using the geographical distances between locations on a map as the input distances to the analyses. The NMDS ordination will re-create the map quite accurately while the PCA ordination will usually not do so.
Some time ago I had the idea of doing this same exercise using a data-display network. Unfortunately, I was beaten to it by Barbara Holland (2013. The rise of statistical phylogenetics. Australian and New Zealand Journal of Statistics 55: 205-220). I will go ahead, anyway, disappointed though I am.
I have chosen the Ukraine as my map. The road distances between 25 of the cities were taken from Ukraine Connections (the same data occur on several other sites, as well).
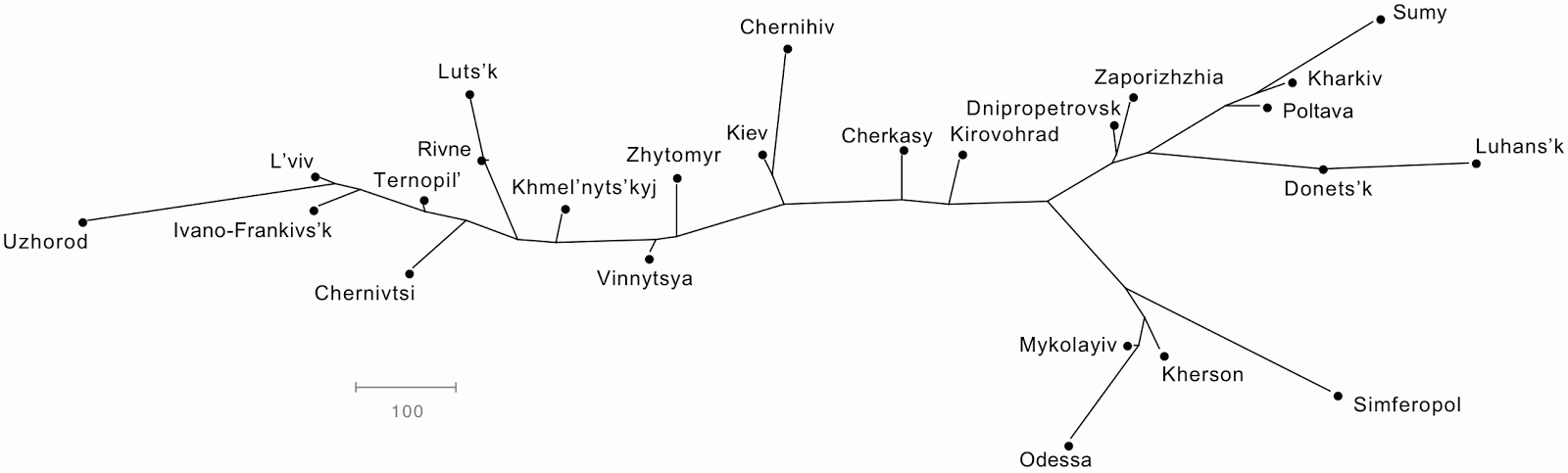
The geographical data were processed in SplitsTree to produce both a Neighbor-Joining tree and a NeighborNet network.
If these techniques are to be effective as data displays, then the positions of the cities in the line graphs should be approximately the same as those in the map. This is, indeed, roughly so, although I had to spend some time manually adjusting the branch angles in the tree (for the best match). The two graphs are more rectangular in overall shape than is the Ukraine, which is somewhat closer to a square, but the relative locations of the points in the graphs do tell you where to look for the cities on the map.
However, the network is the better of the two representations on two grounds. First, the points are constrained to certain locations, and do not need manual adjustment. Second, the network more accurately gives a sense that these are road distances, and there are multiple roads from one city to another — the tree incorrectly implies that there is only one way to get between the cities.
Labels:
EDA,
Map,
Splits graph
Wednesday, October 1, 2014
A fundamental limitation of pedigrees and networks but not trees
It would be nice to think that genealogical history can be reconstructed with ease. However, this is known not to be so. In particular, being able to reconstruct an overall history from a collection of sub-histories, which can thought of as the "building blocks", is not necessarily guaranteed.
That is, even given a complete collection of all of the sub-histories it is not necessarily possible to reconstruct a unique overall history. In other words, there can be pairs of graphs that do not represent the same evolutionary histories, but still display exactly the same collection of building blocks. ("Display" means roughly that a building block can be obtained by simply deleting some of the edges and vertices in the graph.) Mathematically, the sub-histories do not determine (or encode) the history.
For example, it is known that pedigrees cannot necessarily be reconstructed from a collection of all of the sub-pedigrees (Thatte 2008). Pedigrees are the traditional "family trees" showing the ancestry of individuals. Pedigrees differ from phylogenies in that all of the individuals have two parents (rather than possibly having a single immediate ancestor) and there are probably multiple roots (unless there is considerable inbreeding).
Phylogenetic trees, on the other hand can be uniquely reconstructed from a collection of all of the possible sub-trees (see Dress et al. 2012). This is one of the things that makes trees valuable as a phylogenetic model — it is theoretically possible to collect enough information to construct a unique phylogenetic tree.
Rooted phylogenetic networks do not, however, share this property. For some time it has been known that networks cannot necessarily be built from their building blocks, whether those blocks are rooted trees (Willson 2011) or triplets (= rooted 3-taxon trees) or clusters (= rooted sub-trees = clades) (Gambette and Huber 2012).
This is illustrated in the next figure (adapted from Huber et al.), which shows two networks at the top and below that the four trees that are displayed by both of them (by deleting one of each pair of incoming edges at the two reticulation nodes). Given these four trees we cannot reconstruct a unique network, and yet they are the only four trees associated with either network.
To make matters worse, Huber et al. (in press) have now revealed that we can't reconstruct rooted phylogenetic networks even from sub-networks. To do this they show that networks cannot necessarily be built from trinets (= rooted 3-taxon networks). Certain types of networks (e.g. level-1, level-2, tree-child) can be reconstructed (van Iersel and Moulton 2014), but Huber et al. show the example in the second figure, which shows two networks at the top and below that the four trinets that are displayed by both of them. Given these four trinets we cannot reconstruct a unique network, and yet they are the only four trinets associated with either network.
This means that "even if all of the building blocks for some reticulate evolutionary history were to be taken as the input for any given network building method, the method might still output an incorrect history." The best analogy here is Humpty Dumpty — even given all of the pieces, we literally might not be able to put him back together again. We could if he is a rooted tree, but we cannot guarantee it if he is a rooted network or pedigree.
This may not matter in practice, given that we don't yet know the circumstances under which it is possible to uniquely reconstruct networks, but it does mean that we acquire a certain degree of uncertainty as we move from "tree thinking" to "network thinking".
References
Dress A, Huber KT, Koolen J, Moulton V, Spillner A (2012) Basic Phylogenetic Combinatorics. Cambridge Uni Press.
Gambette P, Huber K (2012) On encodings of phylogenetic networks of bounded level. Journal of Mathematical Biology 65: 157-180.
Huber KT, van Iersel L, Moulton V, Wu T (in press) How much information is needed to infer reticulate evolutionary histories? Systematic Biology
van Iersel L, Moulton V (2014) Trinets encode tree-child and level-2 phylogenetic networks. Journal of Mathematical Biology 68: 1707-1729.
Thatte BD (2008) Combinatorics of pedigrees i: counterexamples to a reconstruction problem. SIAM Journal of Discrete Mathematics 22: 961-970.
Willson SJ (2011) Regular networks can be uniquely constructed from their trees. IEEE/ACM Transactions on Computational Biology and Bioinformatics 8: 785-796.
Labels:
Genealogy,
Hybridization network,
Pedigree,
Phylogeny
Subscribe to:
Posts (Atom)