The general public and the general media often have no idea what biologists mean by the work "evolution". The word has two possible meanings, and they usually pick the wrong one. Niles Eldredge tried to clarify the situation by referring to them:
- transformational evolution — the change in a group of objects resulting from a change in each object (often attributed to Lamarck)
- variational evolution - the change in a group of objects resulting from a change in the proportion of different types of objects (usually attributed to Darwin).
So, it was with some trepidation that I looked at an article in Scientific American called Chain letters and evolutionary histories (by Charles H. Bennett, Ming Li and Bin Ma. June 2003, pp. 76-81). It was subtitled: "A study of chain letters shows how to infer the family tree of anything that evolves over time, from biological genomes to languages to plagiarized schoolwork."
The "taxa" in their study consist of 33 different chain letters, collected during the period 1980–1995 (8 other letters were excluded), covering the diversity of chain letters as they existed before internet spam became widespread. These letters can be viewed on the Chain Letters Home Page.
The main issue with this study is that there are no clearly defined characters, from which the phylogeny could be constructed. The authors therefore resort to creating a pairwise distance matrix, among the taxa, in a manner (compression) that I have criticized before (Non-model distances in phylogenetics). I have also discussed previous examples where this approach has been used, notably: Phylogenetics of computer viruses? Multimedia phylogeny?
The essential problem, as I see it, is that without a model of character change there is no reliable way to separate phylogenetic information from any other type of information. That is, phylogenetic similarity is a special type of similarity. It is based on the idea of shared derived character states, as these are the only things that are informative about a phylogeny.
Compression, on the other hand, is a general sort of similarity, based on the idea of information complexity. This presumably will contain some useful phylogenetic information, but it will also contain a lot of irrelevance — for example, shared ancestral character states, which are uninformative at best and positively misleading at worst.
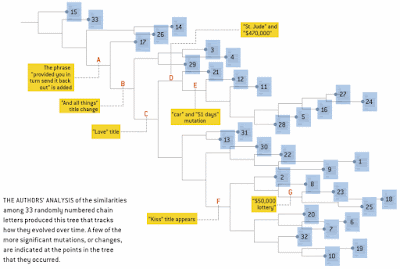
So, the authors can easily produce an unrooted tree from their similarity matrix, which they then proceed to root at one of the letters that they collected early on in their study. This tree is shown here.
However, whether this diagram represents a phylogeny is unknown.
Nevertheless, that does not stop us using an unrooted phylogenetic network as a form of exploratory data analysis, as we have done so often in this blog. This is not intended to produce a rooted evolutionary history, but instead merely to summarize the multivariate information in a comprehensible (and informative) manner. This might indicate whether we are likely to be able to reconstruct the phylogeny In this case, I have used a NeighborNet to display the similarity matrix, as shown next.
It is easy to see that the relationships among the letters are not particularly tree-like. Moreover, the long terminal edges emphasize that much of the complexity information is not shared among the letters, while the shard information is distinctly net-like. So, a simple "phylogenetic tree" (as shown above) is not likely to be representative of the actual evolutionary history.
However, there are actually a few reasonably well-defined groups among the taxa — one at the top. one at the right, and several at the bottom of the network. There are also letters of uncertain affinity, such as L2, L23, L13 and L31. These may reflect phylogenetic history, even though that history is hard to untangle.
Finally, it is worth noting that the history of chain letters, dating back to the 1800s, is discussed in detail by Daniel W. VanArsdale at his Chain Letter Evolution web pages.



No comments:
Post a Comment