This is a guest blog post, following on from his
previous post, by:
Centre des Recherches Linguistiques sur l'Asie Orientale, Paris, France
Introduction
All languages constantly change. Words are lost when speakers cease to use them, new words are gained when new concepts evolve, and even the pronunciation of the words changes slightly over time. Slight modifications that can barely be noticed during a person's lifetime sum up to great changes in the system of a language over centuries. When the speakers of a language diverge, their speech keeps on changing independently in the two communities, and at a certain point of time the independent changes are so great that they can no longer communicate with each other — what was one language has become two.
Demonstrating that two languages once were one is one of the major tasks of
historical linguistics. If no written documents of the ancestral language exist, one has to rely on specific techniques for
linguistic reconstruction (see the examples in this
previous post). These techniques require us to first identify those words in the descendant languages that presumably go back to a common word form in the ancestral language. In identifying these words, we infer
historical relations between them. The most fundamental historical relation between words is the relation of common descent. However, similarly to evolutionary biology, where homology can be further subdivided into the more specific relations of orthology, paralogy, and xenology, more specific fundamental historical relations between words can be defined for historical linguistics, depending on the underlying evolutionary scenario.
Homology and Cognacy in Linguistics and Biology
In evolutionary biology there is a rather rich terminological framework describing fundamental historical relations between genes and morphological characters. Discussions regarding the epistemological and ontological aspects of these relations are still ongoing (see the overview in Koonin 2005, but also this
recent post by David). Linguists, in contrast, have rarely addressed these questions directly. They rather assumed that the fundamental historical relations between words are more or less self-evident, with only few counter-examples, which were largely ignored in the literature (Arapov and Xerc 1974; Holzer 1996; Katičić 1966). As a result, our traditional terminology to describe the fundamental historical relations between words is very imprecise and often leads to confusion, especially when it comes to computational applications that are based on software originally developed for applications in evolutionary biology.
As an example, consider the fundamental concept of
homology in evolutionary biology. According to Koonin (2005: 311), it "designates a relationship of common descent between any entities, without further specification of the evolutionary scenario". The terms orthology, paralogy, and xenology are used to address more specific relations.
Orthology refers to "genes related via speciation" (Koonin 2005: 311); that is, genes related via direct descent.
Paralogy refers to "genes related via duplication" (ibid.); that is, genes related via indirect descent.
Xenology, a notion which was introduced by Gray and Fitch (1983), refers to genes "whose history, since their common ancestor, involves an interspecies (horizontal) transfer of the genetic material for at least one of those characters" (Fitch 2000: 229); i.e. to genes related via descent involving lateral transfer.
In historical linguistics, the only relation that is explicitly defined is
cognacy (also called
cognation). Cognacy usually refers to words related via “descent from a common ancestor” (Trask 2000: 63), and it is strictly distinguished from descent involving lateral transfer (
borrowing). The term
cognacy itself, however, covers both
direct and
indirect descent. Hence, traditionally, German
Zahn 'tooth' is cognate with English
tooth, and German
selig 'blessed' with English
silly, and German
Geburt 'birth' with English
birth, although the historical processes that shaped the present appearance of these three word pairs are quite different. Apart from the sound shape,
Zahn and
tooth have regularly developed from Proto-Germanic
*tanθ-;
selig and
silly both go back to Proto-Germanic
*sæli- 'happy', but the meaning of the English word has changed greatly;
Geburt and
birth stem from Proto-Germanic
*ga-burdi-, but the English word has lost the prefix as a result of specific morphological processes during the development of the English language (all examples follow Kluge and Seebold 2002, with modifications for the pronunciation of Proto-Germanic). Thus, of the three examples of cognate words given, only the first would qualify as having evolved by
direct inheritance, while the inheritance of the latter two could be labelled as indirect, involving processes which are largely language-specific and irregular, such as meaning shift and morpheme loss. Trask (2000: 234) suggests the term
oblique cognacy to label these cases of indirect inheritance, but this term seems to be rarely used in historical linguistics; and at least in the mainstream literature of historical linguistics I could not find even a single instance where the term was employed (apart from the passage by Trask).
In the table above (with modifications taken from List 2014: 39), I have tried to contrast the terminology used in evolutionary biology and historical linguistics by comparing to which degree they reflect fundamental historical relations between words or genes. Here,
common descent is treated as a basic relation which can be further subdivided into relations of
direct common descent,
indirect common descent, and
common descent involving lateral transfer. As one can easily see, historical linguistics lacks proper terms for at least half of the relations, offering no exact counterparts for
homology,
orthology, and
xenology in evolutionary biology.
Cognacy in historical linguistics is often deemed to be identical with
homology in evolutionary biology, but this is only true if one ignores common descent involving lateral transfer. One may argue that the notion of xenology is not unknown to linguists, since the borrowing of words is a very common phenomenon in language history. However, the specific relation which is termed
xenology in biology has no direct counterpart in historical linguistics: the term
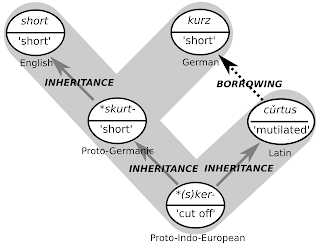
borrowing refers to a distinct process, not a relation resulting from the process. There is no common term in historical linguistics which addresses the specific relation between such words as German
kurz 'short' and English
short. These words are not cognate, since the German word has been borrowed from Latin
cŭrtus 'mutilated' (Kluge and Seebold 2002). They share, however, a common history, since Latin
cŭrtus and English
short both (may) go back to Proto-Indo-European
*(s)ker- 'cut off' (Vaan 2008: 158). The specific history behind these relations is illustrated in the following figure.
A specific advantage of the biological notion of homology as a basic relation covering
any kind of historical relatedness, compared to the linguistic notion of cognacy as a basic relation covering
direct and indirect common descent, is that the former is much more realistic regarding the
epistemological limits of historical research. Up to a certain point, it can be fairly reliably demonstrated that the basic entities in the respective disciplines (words, genes, or morphological characters) share a common history. Demonstrating that more detailed relations hold, however, is often much harder. The strict notion of cognacy has forced linguists to set goals for their discipline which may often be far too ambitious to achieve. We need to adjust our terminology accordingly and bring our goals into balance with the epistemological limits of our discipline. In order to do so, I have proposed to refine our current terminology in historical linguistics to the schema shown in the table below (with modifications taken from List 2014: 44):
Fifty Shades of Cognacy
In a recent
blog post, David pointed to the relative character of homology in evolutionary biology in emphasizing that it "only applies locally, to any one level of the hierarchy of character generalization". Recalling his example of bat wings compared to bird wings, which are homologous when comparing them as
forelimbs but who are analogous when comparing them as
wings, we can find similar examples in historical linguistics.
If we consider words for 'to give' in the four Romance languages Portuguese, Spanish, Provencal and French, then we can state that both Portuguese
dar and Spanish
dar are homologous, as are Provencal
douna and French
donner. The former pair go back to the Latin word
dare 'to give', and the latter pair go back to the Latin word
donare 'to gift (give as a present)'. In those times when Latin was commonly spoken, both
dare and
donare were clearly separated words denoting clearly separated contexts and being used in clearly separated contexts. The verb
donare itself was derived from Latin
donum 'present, gift'. Similarly to English where nouns can be easily used as verbs, Latin allowed for specific morphological processes. In contrast to English, however, these processes required that the form of the noun was modified (compare English
gift vs.
to gift with Latin
donum vs.
donare).
What the ancient Romans (who spoke Latin as their native tongue) were not aware of is that Latin
donum 'gift' and Latin
dare 'to give' themselve go back to a common word form. This was no longer evident in Latin, but it was in Proto-Indo-European, the ancestor of the Latin language. Thus, Latin
dare goes back to Proto-Indo-European
*deh3- 'to give', and Latin
donum goes back to Proto-Indo-European
*deh3-no- 'that which is given (the gift)' (Meiser 1999; what is written as
*h3 in this context was probably pronounced as [x] or [h]). The word form
*deh3-no- is a regular derivation from
*deh3-, so at the Indo-European level both forms are homologous, since one is derived from the other. That means, in turn, that Latin
dare and
donum are also homologs, since they are the residual forms of the two homologous words in Proto-Indo-European. And since Latin
donare is a regular derivation of
donum, this means, again, that Latin
dare and
donare are also homologous, as are the words in the four descendant languages, Portuguese
dar, Spanish
dar, Provencal
douna, and French
donner. Depending on the time depth we apply, we will arrive at different homology decisions. I have tried to depict the complex history of the words in the following figure:
Judging from the treatment in linguistic databases, many scholars do not regard these different "shades of homology" as a real problem. In most cases, scholars use a "lumping approach" and label as cognates all words that go back to a common root, no matter how far that root goes back in time (compare, for example, the cognate labeling for reflexes of Proto-Indo-European
*deh3- in the
IELex).
Importantly, this labeling practice, however, may be contrary to the models that are used to analyze the data afterwards. All computational analyses model language evolution as a process of word gain and word loss. The words for the analyses are sampled from an initial set of concepts (such as 'give', 'hand', 'foot', 'stone', etc.) which are translated into the languages under investigation. If we did not know about the deeper history of Latin
dare and
donare, we would assume a regular process of language evolution here: at some point, the speakers of Gallo-Romance would cease to use the word dare to express the meaning 'to give' and use the word
donare instead, while the speakers of Ibero-Romance would keep on using the word
dare. This well-known process of
lexical replacement (illustrated in the graphic below), which may provide strong phylogenetic signals, is lost in the current encoding practice where all four words are treated as homologs. Our current practice of cognate coding masks vital processes of language change.
Outlook
Historical linguistics needs a more serious analysis of the fundamental processes of language change and the fundamental historical relations resulting from these processes. In the last two decades a large arsenal of quantitative methods has been introduced in historical linguistics. The majority of these methods come from evolutionary biology. While we have quickly learned to adapt and apply these methods to address questions of language classification and language evolution, we have forgotten to ask whether the processes these methods are supposed to model actually coincide with the fundamental processes of language evolution. Apart from adapting only the methods from evolutionary biology, we should consider also adapting the habit of having deeper discussions regarding the very basics of our methodology.
References
Arapov MV, Xerc MM (1974) Математические методы в исторической лингвистике [Mathematical methods in historical linguistics]. Moscow: Nauka. German translation: Arapov, M. V. and M. M. Cherc (1983).
Mathematische Methoden in der historischen Linguistik. Trans. by R. Köhler and P. Schmidt. Bochum: Brockmeyer.
Fitch WM (2000) Homology: a personal view on some of the problems.
Trends in Genetics 16.5, 227-231.
Gray GS, Fitch WM (1983) Evolution of antibiotic resistance genes: the DNA sequence of a kanamycin resistance gene from
Staphylococcus aureus.
Molecular Biology and Evolution 1.1, 57-66.
Holzer G (1996)
Das Erschließen unbelegter Sprachen. Zu den theoretischen Grundlagen der genetischen Linguistik. Frankfurt am Main: Lang
Katičić R (1966) Modellbegriffe in der vergleichenden Sprachwissenschaft.
Kratylos 11, 49-67.
Kluge F, Seebold E (2002)
Etymologisches Wörterbuch der deutschen Sprache. 24th ed. Berlin: de Gruyter.
List J-M (2014)
Sequence Comparison in Historical Linguistics. Düsseldorf: Düsseldorf University Press.
Meiser G (1999)
Historische Laut- und Formenlehre der lateinischen Sprache. Wissenschaftliche Buchgesellschaft: Darmstadt.
Trask RL (2000)
The Dictionary of Historical and Comparative Linguistics. Edinburgh: Edinburgh University Press.
Vaan M (2008)
Etymological Dictionary of Latin and the Other Italic Languages. Leiden and Boston: Brill.