In a recently published pre-print on PaleorXiv, Roland Sookias makes a point for distinguishing between parallelism, ie. shared inherited traits that can be found in some but not all of the offspring of a common ancestor, and convergences in a strict sense, involving similar traits that are not homologous. The former is also known as homoiology, a term Sookias attributes to Ludwig Plate.
As a geneticist working mostly at the tips of the Tree of Plant Life, I'm quite familiar with the (pre-Hennigian) concept: we much more often than not lack Hennig's 'synapomorphies', ie. shared, derived traits exclusive to an evolutionary lineage. But we have many highly diagnostic characters suites including 'shared apomorphies' (I think that the angiosperm phylogeneticist Jim Doyle coined the term) that collect the same species or higher taxa, eg. groups of taxa that also form highly supported clades in molecular trees, but are not exclusive. In every plant group you can additionally observe that certain traits are exclusive to some members of one lineage, because the lineage has the genetic-physiological prerequisites to express these traits, while their sister lineages or distant relatives lack this potential. Epigenetics deals with tendencies to express a trait in response to the environment without even changing the genetic code.
If you look close enough, you can find such patterns even at the molecular level.
 |
| Molecular evolution of the 5' half of the ITS1 in beeches. Each sequence motif is assigned a state (Ax, Bx etc; x = 0 represents the ancestral state, x > 0 are derived states) and evolution involves usually the gain ("+") or loss ("-") of sequence motifs including some potential genetic homoiologies (see here for context and references). |
However, it has apparently been ignored by my fellow paleontologists: Sookias' wants to discuss "the neglected concept of homoiology ... in the context of palaeontological phylogenetic methods". Paleontological phylogenetic methods are, of course, tree inferences, and the idea is that recognition of homoiologies can be a means of establishing node support or to "help to choose between equally parsimonious or likely trees". He provides an R function "to calculate two measures for a given tree and matrix: (a) the potential support for clades based on potential homoiologies; and (b) the fit of the tree to all states given the concept of homoiology".
Sookias provides a nice and conscise introduction to the problem with some examples, and makes the connection to linguistics (see also Mattis' and my post on the Chinese dialects continuum: How languages lose body parts); so, give the short paper a read. Like all paleontological literature it is strongly influenced by cladistic views, such as that life is monophyletic, and it revolves around the central theme how to get better supported trees.
My inner geneticist has a principal problem with such a goal, because there has (to my knowledge) not been a single morphology-based tree that was fully congruent to a molecular tree with sufficient taxon and gene sampling, which applies also to the real-world data example that Sookias chose (as we will see).
My inner paleontologists also knows that there are highly diagnostic morphs in the fossil record, but diagnostic character suites and morphs reflect as many paraphyla as monophyla. He also knows that the fossil record, provided you find the right fossil from the right time, may alter your perspective on ancestral and derived character states.
 |
| An inferred tree (see this post). Given the inferred tree (quasi-dated tree), we would assume that star shapes are primitive (a symplesiomorphy) within the Pointish lineage, and possibly 10-tipped stars; and conclude that the Tenstars are paraphyletic. Greenish is clearly ancestral (a Pointish symplesiomorphy), and bluish derived (a Polygonia synapomorphy). |
So, why should we aim to get more resolved, better supported, morphology-based trees? Any such tree will inevitably include wrong branches!
I argue that, instead, we should just explore the signal in our data matrices using networks. Any potential tree is included in a network. But networks are more comprehensive because they provide not only a single tree but alternative, competing trees. By visualizing the alternatives, we can discern between mere convergence (random similarity), homoiology (parallelism, convergence related to descent), symplesiomorphy (shared, lineage-consistent primitive traits) and synapomorphy (lineage-unique and consistent shared derived traits), which can be very tricky with just a tree. Thus, we can try to evaluate which evolutionary scenario best explains all our data.
Compatibility
The basic problem when using morphological and such-like data sets to infer phylogenies is that most of the scored characters are, to some degree, incompatible with the true tree, ie. the actual evolutionary pathways.
Let's take a hypothetical evolution (no reticulations), in which the x-axis represents the morphological diversification and the y-axis time.
As in real-world data, sister taxa (eg. Species A and B) may have different levels of morphological derivation compared to their common ancestor(s). This leads us to this unrooted true tree in which the branch lengths are proportional to the real (above) amount of change.
 |
| Unrooted representation of the above evolution. All commonly used tree inferences infer unrooted trees. |
The only characters providing a taxon bipartition that is fully compatible with the true tree are Hennig's 'synapomorphies':
 |
| Clade A–D shares a unique, derived trait. The character split is fully compatible with the true tree. |
Next come Hennig's 'symplesiomorphies' (Sookias' R-script discards them):
Homoiologies / parallelisms can be partly compatible:
Note that without a molecular backbone tree, it may be impossible to distinguish homoiologies from symplesiomorphies – whether a trait will be resolved as either the one or the other in a tree depends solely on its frequency and distribution across the subtree, and the situation in outgroups.
 |
| Purple is the plesiomorphy of the ingroup, blue the homoiology found in members of Clade A–F, evolved twice |
Doyle's 'shared apomorphies' are locally compatible:
Convergence in a strict sense, ie. superficial or random similarity, is incompatible with the true tree:
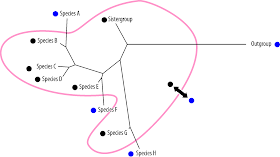 |
| Blue is a randomly distributed derived state found in all longer-branched taxa. |
A tree-incompatible signal is, naturally, best handled using a network and not by forcing it into a single tree. Unless, of course, we have a sensible molecular tree and can go for total evidence approaches assuming the molecular tree reflects the true tree.
PS: Also, in molecular data the true tree incompatible characters may outnumber the compatible ones, but there we have many more characters and (usually but not always) a lot that are not filtered by negative or positive selection. Our stochastic molecular models are for sure never accurate enough to model molecular evolution for our sequences, but apparently precise enough for most applications. Even before next generation sequencing and big data, molecular phylogenies outshined morphological phylogenies, something that paleontologists cannot afford to ignore any more — not because the data are much better (to infer evolution) but because the patterns and processes are much less complex.
Sookias' data example, crocodiles and relatives
The supplement of Sookias' paper includes a morphological character matrix for crocodilians and the resulting molecular tree for the group. Here's Sookias' fig. 3 ,using these data to make his point for how to select the better-fitting tree using homoiology recognition:
Now, the unsolved problem is: if we don't have a molecular tree, how can we possibly know 0 is a homoiology and not a symplesiomorphy, 1 not a reversal (scenario B) or likely convergence (scenario C), hence, B should be preferred over C (the legend has a little typo, cf. Sookias 2019, p. 3, l. 34)?
The matrix provided as the example is not the best one to make this point. Sookias' script, when stringently eliminating potential symplesiomorphies, identifies, using the molecular tree as input, one potential homoiology for the Crocodylinae, five for their larger clade (including Gavialis and Tomistoma), and one for the alligators' larger clade in a matrix with 117 characters. Less than 10% can hardly be a game-changer.
What the morpho-data shows
Furthermore, the morphological matrix will give us a single most-parsimonious tree (MPT, using PAUP*'s Branch-and-Bound algorithm), not two or more equally parsimonious alternatives that we need to weigh against each other.
Some of the red branches are supported by pseudo-synapmorphies, which, on the background of the molecular tree, are potential homoiologies for the comprising clade, however, interpreted as symplesiomorphies by Sookias' script (provided the molecular branch-lengths are sufficient, they might be recognized when using a probabilistic framework to infer the ancestral states).
Not a good example for Sookias goal, but the matrix shows the limitations of trees when it comes to morphological differentiation. Here's the distance-based, 2-dimensional network for the morphological data:
 |
| A Neigbor-net based on Sookias' morphological matrix. The arrow indicates the position of the assumed root. |
The signal from the morphological matrix is quite tree-like, and the structure of the left part of the network is synonymous to that of the single MPT (and the molecular tree). On the right-hand side, we find more complexity than we would expect from the single MPT. The data signal is not trivial regarding the position of the root as inferred by Bernissartia; and nor is the placement of Gavialis and Tomistoma (pink edge bundles), two genera producing a very prominent box-like structure. Called by cladists a "phenetic" approach, the distance-based network is nonetheless straightforward regarding the identification of monophyletic groups (green) and potential monophyletic groups (yellow) (the latter always include the particular alternative seen in the single MPT as well, in case of the pink box, also the molecular alternative). The light green monophylum is a necessary consequence of the prior knowledge about the position of the root, and the likely monophyly of Alligator and its relatives (the tree-like subgraph with long internal branches = lots of uniquely shared traits, including potential synapmorphies).
 |
| Potential synapomorphies that can be inferred from the morpho-matrix alone by mapping the states onto the network. Red, homoiologies reconstructed as synapomorphies ('pseudo-synapomorphies') and (except for one) excluded as potential symplesiomorphies by Sookias' test run of his script (strict and relaxed cut-off). |
The network provides more information than can be extracted from the MPT: one Crocodylus is significantly closer to the Osteolaemus (the neighborhood defined by the light blue edge bundle, see Sookias' fig. 3A). Crocodylus, however, is likely monophyletic, being generally very similar; and the third genus, Mecitops, is closely linked to (all of) them (neighbourhood defined by the dark blue edge). An inclusive common origin (including the third genus, Mecistops) is – just based on morphology and without using a "phylogenetic" tree inference – beyond question, even though we lack syn- or shared apomorphies (short corresponding edge bundle): Mecistops is obviously closely related to Crocodylus, and Osteolaemus is related to part of the latter, so it's not a bad hypothesis that all three are descendants of the same common ancestor, and that Tomistoma (and Gavialis) branched off the lineage before the Crocodylinae radiated. The only alternative explanation would be that the Crocodylinae show the primitive morphs of the entire lineage, and that the position of Tomistoma and Gavialis is affected by long-branch (-edge) attraction (however, if that is the case then we should have found a Tomistoma-Gavialis clade in the MPT — parsimony will always get it wrong in the Felsenstein zone)
The main flaw
But, any morphology-based alternative using this data matrix is not fully compatible with the molecular tree, which places Mecitops and Osteolaemus as sister to Crocodylus. Here's the consensus network based on 10,000 boostrap pseudoreplicate BioNJ trees inferred from the morpho-matrix, highlighting the support for splits compatible with the molecular tree (green) and their competing, partly incongruent (red edge bundles) alternatives (I do the information transfer manually, but those with R-scripting skills can use the functions in the phangorn library; Schliep et al., MEE, 2017; see also David's post):
Assuming the molecular tree is correct, the above reconstruction means that Osteolaemus is morphologically more derived, and hence placed as sister, while Mecitops and Crocodylus retain more primitive character states, and hence lacks discriminatiory derived traits — a sort of local ingroup-outgroup long-branch attraction (or 'short-branch culling').
We can also hypothesize that the initial radiation was fast, because the Mecitops-Osteolaemus ancestor did not accumulate a single, unique, discriminating character trait.
Excess of shared derived, pseudo-synapomorphic traits is the reason Tomistoma is not resolved as sister of Gavialis in the MPT — the molecular Gavialis-Tomistoma clade is represented by a morphological grade.
And the homoiologies identified using the molecular tree as input cannot put things right. They are just partly compatible with unproblematic splits, ie. the larger clade including Alligator (character #7), the larger clade including Crocodylinae (#1, #18, #73, #74, #117) or exclusive to the Crocodylinae (#66)
 |
| Character mapping of the molecular-inferred homoiologies. The lush green splits represent the molecular splits. |
However, if we are ignorant of the molecular tree, we would have to assume that Mecitops is the sister to Crocodylus, and that some of their shared traits not found in Osteolaemus are shared apomorphies (if occurring outside the clade and in the sister clade) or even synapomorphies (if exclusive for Mecitops + Crocodylus), while only those shared by Osteolaemus and C. porosus (#66) can be homoiologies. We also would have no reason to challenge the Gavialis-Tomistoma grade, until we infer networks.
This is the main flaw of Sookias' idea. To identify homoiologies, we need the same prerequisite as for any of Hennig's concepts: we need to know the true tree. If we use the inferred tree based on the same data that we want to weight (here: use homoiologies for decision making or means of node support), then we propagate first-level errors; apply circular reasoning. Such as the red-marked pseudo-synapomorphies in the network above; vice versa, all actual (molecular-wise) synapomorphies supporting the molecular Gavialis-Tomistoma clade (dark purple split) would be reconstructed as homoiologies or symplesiomorphies based on the morpho-based single MPT (or morpho-based NJ tree, or probabilistic tree).
And if we have an independent molecular tree, it will make the decision on the fly: putative synapormorphies are the traits that are fully compatible, symplesiomorphies, homoiologies and shared apomorphies are decreasingly compatible, and random convergences are incompatible with the molecular tree.
It is not homoiology but tree-incompatible signal that is neglected in phylogenetics
Sookias points out: "In inference of phylogeny by parsimony, an occurrence of a character state in a part of a tree separated from it by another state is considered simply a homoplasy, and a tree where the occurrences are nearer or further from one another is not more or less parsimonious ... a tree where the 15 occurrences are nearer or further from one another is not more or less parsimonious". In principle, this is true, but has little consequence in application.
We, usually without realizing it, make frequent use of the discriminating power of potential homoiologies. See the example above, but also when, eg., placing fossils in a molecular framework or do post-inference character weighting. In these cases, homoiologies (and symplesiomorphies) will stabilize the inference and increase support. For better and worse:
- Better, because homoiologies will ensure that the fossil is placed in the right molecular-based subtree, and can compensate for the lack of synapomorphies. Imagine an extinct fossil sibling lineage showing only homoiologies shared by Osteolaemus and C. porosus. Using tree-based optimization (eg. RAxML's 'evolutionary placement algorithm'), it would be placed close to the Crocodylinae ancestor, likely next to Osteolaemus. Using a Neighbor-net, it would be placed between Osteolaemus and C. porosus. Either way, the homoiologies would ensure it is nested within the Crocodylinae.
- Post-inference character weighting, as implemented in eg. TNT, will downweight inferred convergences (ie. higher homoplasy, more stochastically distributed across the tree) more than putative homoiologies (ie. less homoplastic since confined to a single subtree). This can be better or worse. How do we avoid what happened for the crocodiles that homoiologies are not recognized as such but support (somewhat) misleading clades (act as synapomorphies)? Clades are commonly interpreted as a sufficient criterion to determine monophyly; however, they are not even a necessary one: taxa can be part of a monophyletic group despite not forming an inclusive subtree (ie. clade in a rooted tree) such as the genus Caiman or Gavialia-Tomistoma.
One thing is also obvious from the crocodile example: if we have enough signal in the morphological data, then we may get one or another thing wrong and, in some cases, may not be able to decide between one or another alternative. However, overall, the morphological differentiation pretty well captures what the genes provide us as the best approximation of the true tree. Even when the matrix includes very few potential synapomorphies and clear homoiologies but a lot of shared apomorphies, most of which were convergently evolved in parts of both major clades.
At least, this will be so when we analyze the data using networks and not just trees (compare the single MPT to the networks).
Using the alternative evolutionary scenarios provided by the networks, we can then look back into our data (see the maps above), to see what may be a homoiology, a symplesiomorphy (very useful for deciding between evolutionary scenarios, as well) or a synapomorphy. The phangorn library (used for Sookias' script) has now network functionality and allows transferring information between trees and networks. An R-affine person may be able to extract lists of potential (partly competing) synapomorphies, symplesiomorphies, and homoiologies directly from the network showing all possible or the most likely trees.
And then use this information to eg. place fossils in a phylogenetic context, or reconstruct evolutionary trends in extinct groups of organisms — reconstruction of evolutionary trends in extant organisms should always rely on morphological data analyzed in a molecular-phylogenetic framework.
Data
A NEXUS-version of Sookias' test matrix (slightly annotated for Mesquite, simple version for PAUP*), tree- and distance matrix files have been added to my figshare collection of morphological matrices.













No comments:
Post a Comment